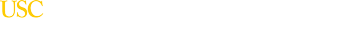Publications and Events
Publications
La Rocca M, Garner R, Amoroso N, Lutkenhoff ES, Monti MM, Vespa P, Toga AW, Duncan D. . Multiplex Networks to Characterize Seizure Development in Traumatic Brain Injury Patients. Front Neurosci. ;14:591662. doi: 10.3389/fnins.2020.591662. PMID: 33328863; PMCID: PMC7734183. . 2021 Nov 30.
Faghihpirayesh R, Ruf S, La Rocca M, Garner R, Vespa P, Erdogmus D, Duncan D.. Automatic Detection of EEG Epileptiform Abnormalities in Traumatic Brain Injury using Deep Learning. Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) 2021. . 2021 Nov 1.
Akbar MN, Ruf S, La Rocca M, Garner R, Barisano G, Cua R, Vespa P, Erdogmus D, Duncan D.. esion Normalization and Supervised Learning in Post-traumatic Seizure Classification with Diffusion MRI. International Workshop on Computational Diffusion MRI. . 2021 Oct 1.
https://doi.org/10.1007/9
Lutkenhoff ES, Shrestha V, Ruiz Tejeda J, Real C, McArthur DL, Duncan D, La Rocca M, Garner R, Toga AW, Vespa PM, Monti MM. Early brain biomarkers of post-traumatic seizures: initial report of the multicentre epilepsy bioinformatics study for antiepileptogenic therapy (EpiBioS4Rx) prospective study. Journal of neurology, neurosurgery, and psychiatry. ;91(11):1154-1157. PubMed PMID: 32848013; PubMed Central PMCID: PMC7572. . 2020 Nov 9.
Lutkenhoff ES, Wright MJ, Shrestha V, Real C, McArthur DL, Buitrago-Blanco M, Vespa PM, Monti MM. The subcortical basis of outcome and cognitive impairment in TBI: A longitudinal cohort study. Neurology. PubMed PMID: 32907958; DOI: 10.1212/WNL.0000000000010825. . 2020 Sep 9.
Liu J, Garner R, La Rocca M, Bae EK, Duncan D. . The effects of filtering on high frequency oscillation classification. 2020 IEEE Spring Simulation Conference (SpringSim). doi: 10.22360/SpringSim.2020.MSM.008.. . 2020 May 18.
Cabeen RP, Immonen R, Harris NG, Gröhn O, Smith G, Manninen E, Garner R, Duncan D, Pitkänen A, Toga AW. . A computational diffusion mri framework for biomarker discovery in a rodent model of post-traumatic epileptogenesis. 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI). doi: 10.1109/ISBI45749.2020.9098575.. . 2020 Apr 3.
Kwon CS, Agarwal P, Subramaniam V, Dhamoon M, Mazumdar M, Yeshokumar A, Panov F, Ghatan S, Jetté N.. Readmission after neurosurgical intervention in epilepsy: A nationwide cohort analysis. Epilepsia.;61(1):61-69. PubMed PMID: 31792965; PubMed Central PMCID: PMC7227389; DOI: 10.1111/epi.16401.. . 2020 Jan 8.
Nariai H, Hussain SA, Bernardo D, Motoi H, Sonoda M, Kuroda N, Asano E, Nguyen JC, Elashoff D, Sankar R, Bragin A, Staba RJ, Wu JY. Scalp EEG interictal high frequency oscillations as an objective biomarker of infantile spasms. Clinical neurophysiology : official journal of the International Federation of Clinical Neurophysiology.3;131(11):2527-2536. PubMed PMID: 32927206; DOI: 10.1016/j.clinph.2020.08.013.. . 2020 Jan 6.
Garner R, La Rocca M, Vespa P, Jones N, Monti MM, Toga AW, Duncan D. Imaging biomarkers of posttraumatic epileptogenesis. Epilepsia. 2019 Oct 4.
Santana-Gomez C, Andrade P, Hudson MR, Paananen T, Ciszek R, Smith G, Ali I, Rundle BK, Ndode-Ekane XE, Casillas-Espinosa PM, Immonen R, Puhakka N, Jones N, Brady RD, Perucca P, Shultz SR, Pitkänen A, O'Brien TJ, Staba R. Harmonization of pipeline for detection of HFOs in a rat model of post-traumatic epilepsy in preclinical multicenter study on post-traumatic epileptogenesis. Epilepsy Research. 2019 Oct 2.
Engel J, Pitkanen A. Biomarkers For Epileptogenesis And Its Treatment. Neuropharmacology. 2019 Aug 1.
Li Lin, Bragin A, Staba R, Engel J. Unit firing and oscillations at seizure onset in epileptic rodents. Neurobiology of Disease. 2019 Jul 17.
Agoston DV, Vink R, Helmy A, Risling M, Nelson D, Prins M.. How to Translate Time: The Temporal Aspects of Rodent and Human Pathobiological Processes in Traumatic Brain Injury. Journal of neurotrauma;36(11):1724-1737. PubMed PMID: 30628544; DOI: 10.1089/neu.2018.6261.. . 2019 Jun 6.
Garner R, La Rocca M, Barisano G, Vespa P, Toga AW, Duncan D. A machine learning model to predict seizures susceptibility from resting-state fMRI connectivity. Proceedings: Spring Simulation Conference for the Society for Modeling and Simulation. Neurobiology of Disease. 2019 Apr 11.
Agoston DV and Kamnaksh A. Protein biomarkers of epileptogenicity after traumatic brain injury. Neurobiology of Disease. 2019 Mar 29.
Semple BD, Zamani A, Rayner G, Shultz SR, Jones NC. Affective, neurocognitive and psychosocial disorders associated with traumatic brain injury and post-traumatic epilepsy. Neurobiology of Disease. 2019 Mar 28.
Dadas A, Janigro D. Breakdown of blood brain barrier as a mechanism of post-traumatic epilepsy. Neurobiology of Disease. 2019 Mar 27.
Galanopoulou AS, Engel J, Moshé SL. PREFACE: Antiepileptogenesis following traumatic brain injury. Neurobiology of Disease. 2019 Mar 26.
Brady RD, Casillas-Espinosa PM, Agoston DV, Bertram EH, Kamnaksh A, Semple BD, Shultz SR. Modelling traumatic brain injury and posttraumatic epilepsy in rodents. Neurobiology of Disease. 2019 Mar 22.
Hunter LE, Branch CA, Lipton ML. The neurobiological effects of repetitive head impacts in collision sports. Neurobiology of Disease. 2019 Mar 21.
Vespa PM, Shrestha V, Abend N, Agoston D, Au A, Bell MJ, Bleck TP, Blanco MB, Claassen J, Diaz-Arrastia R, Duncan D, Ellingson B, Foreman B, Gilmore EJ, Hirsch L, Hunn M, Kamnaksh A, McArthur D, Morokoff A, O'Brien T, O'Phelan K, Robertson CL, Rosenthal E, Staba R, Toga A, Willyerd FA, Zimmermann L, Yam E, Martinez S, Real C, Engel J Jr. The epilepsy bioinformatics study for anti-epileptogenic therapy (EpiBioS4Rx) clinical biomarker: Study design and protocol. Neurobiology of Disease. 2019 Mar 20.
Immonen R, Smith G, Brady. Harmonization of pipeline for preclinical multicenter MRI biomarker discovery in a rat model of post-traumatic epileptogenesis. Epilepsy Research. 2019 Feb 15.
Engel J, Bragin A, Staba R. Nonictal EEG biomarkers for diagnosis and treatment. Epilepsy Open. 2018 Dec 17.
Zhang J, Bekkers E, Chen D, Berendschot TTJM, Schouten J, Pluim JPW, Shi Y, Dashtbozorg B, Romeny BMTH.. Reconnection of Interrupted Curvilinear Structures via Cortically Inspired Completion for Ophthalmologic Images. IEEE transactions on bio-medical engineering; 65(5):1151-1165. PubMed PMID: 29683430; PubMed Central PMCID: PMC6880863; DOI: 10.1109/TBME.2017.2787025.. . 2018 May 5.
Zhan L, Jenkins LM, Wolfson OE, GadElkarim JJ, Nocito K, Thompson PM, Ajilore OA, Chung MK, Leow AD. The significance of negative correlations in brain connectivity. The Journal of comparative neurology; 525(15):3251-3265. PubMed PMID: 28675490; PubMed Central PMCID: PMC6625529; DOI: 10.1002/cne.24274.. . 2017 Oct 15.
Faghihpirayesh R, Ruf S, La Rocca M, Garner R, Vespa P, Erdogmus D, Duncan D. Automatic Detection of EEG Epileptiform Abnormalities in Traumatic Brain Injury using Deep Learning. Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) 2021. 2021 Nov 1.
Akbar MN, Ruf S, La Rocca M, Garner R, Barisano G, Cua R, Vespa P, Erdogmus D, Duncan D. Lesion Normalization and Supervised Learning in Post-traumatic Seizure Classification with Diffusion MRI. International Workshop on Computational Diffusion MRI. 2021 Oct 1. https://doi.org/10.1007/978-3-030-87615-9_12
La Rocca M, Garner R, Amoroso N, Lutkenhoff ES, Monti MM, Vespa P, Toga AW, Duncan D. Multiplex Networks to Characterize Seizure Development in Traumatic Brain Injury Patients. Front Neurosci. 2020 Nov 30;14:591662. doi: 10.3389/fnins.2020.591662. PMID: 33328863; PMCID: PMC7734183.
Liu J, Garner R, La Rocca M, Bae EK, Duncan D. The effects of filtering on high frequency oscillation classification. 2020 IEEE Spring Simulation Conference (SpringSim). 2020 May 18. doi: 10.22360/SpringSim.2020.MSM.008.
Cabeen RP, Immonen R, Harris NG, Gröhn O, Smith G, Manninen E, Garner R, Duncan D, Pitkänen A, Toga AW. A computational diffusion mri framework for biomarker discovery in a rodent model of post-traumatic epileptogenesis. 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI). 2020 Apr 3. doi: 10.1109/ISBI45749.2020.9098575.
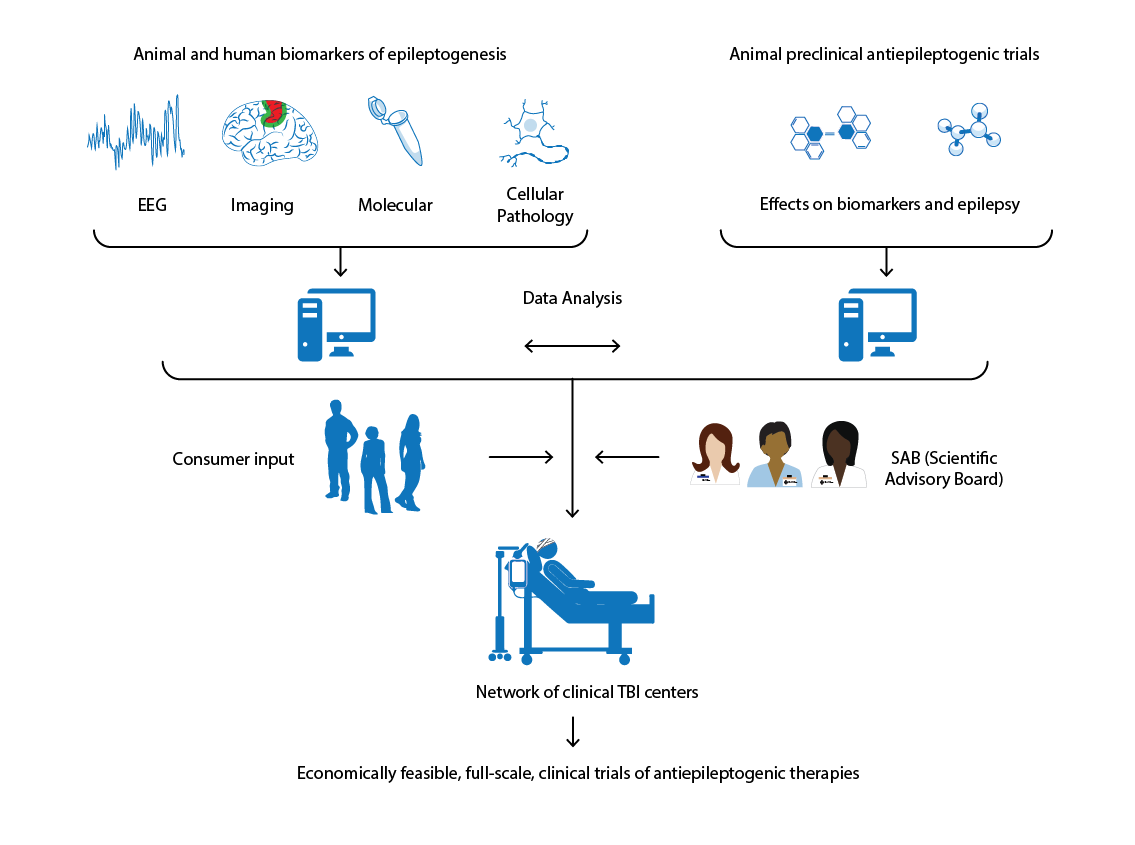
Duncan D, Barisano G, Cabeen R, Sepehrband F, Garner R, Braimah A, Vespa P, Pitkanen A, Law M, Toga AW. Analytic Tools for Post-traumatic Epileptogenesis Biomarker Search in Multimodal Dataset of an Animal Model and Human Patients. Frontiers in Neuroinformatics. 2018;12:86.
Immonen R, Harris NG, Wright D, Johnston L, Manninen E, Smith G, Paydar A, Branch C, Grohn O. Imaging biomarkers of epileptogenecity after traumatic brain injury - Preclinical frontiers. Neurobiology of disease. 2018 Oct 12.
Correa DJ, Kwon CS, Connors S, Fureman B, Whittemore V, Jetté N, Matthern GW, Moshé SL, for the EpiBioS4Rx Public Engagement Core. Applying participatory action research in traumatic brain injury studies to prevent post-traumatic epilepsy. Neurobiology of Disease. 2018 Jul 18.
Saletti PG, Ali I, Casillas-Espinosa PM, Semple BD, Lisgaras C, Moshé SL, Galanopoulou AS. In search of antiepileptogenic treatments for post-traumatic epilepsy. Neurobiology of Disease. 2018 Jun 22.
Perucca P, Smith G, Santana-Gomez C, Bragin A, Staba R. Electrophysiological biomarkers of epileptogenicity after traumatic brain injury. Neurobiology of disease. 2018 Jun 5.
Duncan D, Vespa P, Pitkanen A, Braimah A, Lapinlampi N, Toga AW. Big data sharing and analysis to advance research in post-traumatic epilepsy. Neurobiology of disease. 2018 Jun 1.
Tubi MA, Lutkenhoff E, Blanco MB, McArthur D, Villablanca P, Ellingson B, Diaz-Arrastia R, Van Ness P, Real C, Shrestha V, Engel J. Early seizures and temporal lobe trauma predict post-traumatic epilepsy: a longitudinal study. Neurobiology of disease. 2018 Jun 1.
Pitkänen A, Ndode-Ekane X, Lapinlampi N, Puhakka N. Epilepsy biomarkers–Toward etiology and pathology specificity. Neurobiology of disease. 2018 May 18.
Engel J. Epileptogenesis, traumatic brain injury, and biomarkers. Neurobiology of disease. 2018 Apr 3.
Events
Investigators meeting at UCLA, June 2018
Investigators meeting at NIH, December 2017
Investigators meeting in Houston, TX, December 2016